Links
alphanumeric index
sdf archives
numeric index
pdb files by numeric
sdf files by numeric
Search warning: not yet fully spidered
|
This site is under constant construction
The molecule of the day is: D-myo-Inositol
Inositol is a key molecule in the calcium signaling pathways of the cell. Enzymes
that are involved in Inositol pathways are becoming important targets for cancer
drugs and other therapies. This is a hot molecule!
other names: 1-Diphosinositol pentakisphosphate 148077-18-3 C11174 D-myo-Inositol, 2,3,4,5,6-pentakis(dihydrogen phosphate)1-(trihydrogen diphosphate)
related molecule names: 121010-58-0 1D-myo-Inositol 1,4,5,6-tetrakisphosphate C11555 D-myo-Inositol 1,4,5,6-tetrakisphosphate Inositol 1,4,5
,6-tetrakisphosphate 1D-myo-inositol 1,3,4,6-tetrakisphosphate CPD-505 D-myo-inositol 1,3,4,6-tetrakisphosphate Inositol 1,3,4,6-tetrakis
phosphate 1D-myo-Inositol 1,2,3,4,5,6-hexakisphosphate D-myo-Inositol 1,2,3,4,5,6-hexakisphosphate Inositol 1,2,3,4,5,6-hexaki
sphosphate InsP6 MI-HEXAKISPHOSPHATE 1D-myo-Inositol 1,4,5-trisphosphate 85166-31-0 88269-39-0 D-myo-Inositol 1,4,5-trisphosphate inositol 1,4,5-trisphos 1D-myo-Inositol 1L-myo-Inositol 87-89-8 Bios I C00137 Cyclohexitol D-myo-Inositol
molecule directory: 12652
related molecule directories:
3384
4338
4360
4374
4397
4421
4432
related molecule pdb files:
3384.pdb
4338.pdb
4360.pdb
4374.pdb
4397.pdb
4421.pdb
4432.pdb
|
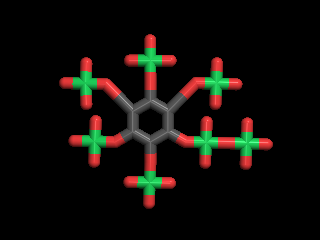
pdb file, 12652.pdb
CAS: 148077-18-3
|
|
Molecule News
updated Tue Aug 19 19:45:09 EDT 2008
Some facts: The Molecules website contains more than
4 million small molecule structure files in pdb format,
and molecular graphics representations. About 50
million molecules are still in the pipe, and they are expected
to appear here over the course of the next few weeks and months.
The pdb format is readable by common FOSS molecule viewer software,
such as RasMol and PyMOL.
In due course, we plan to provide high quality structures via
energy minimization refinement, and additional resources.
Molecules@gnu-darwin.org is founded
in the spirit of free software, open source, and public access.
It is hoped that access to these files will be a wonderful community
resource for science education, research, and entertainment as well.
We are looking for investment or funding to expedite and expand this work, and lead the field,
with an eye towards an advanced, complete, synthetic, structural, and informatical bioorganome.
Meanwhile, the site is already an exceptional lab resource,
and molecular catalog, providing the means and building blocks
towards additional novel structures. We aim to be the best.
The structural biology, protein crystallography, and molecular graphics talent that is
building the Molecules archive is available to work for you in
a contract or consulting arrangement. Wide-ranging expertise is available.
Molecules@gnu-darwin.org is built entirely with
FOSS,
free and open source
software, GNU-Darwin OS, and it is under
the aegis of
The GNU-Darwin Distribution.
Here is a link to the
Distribution
résumé.
Our founder is an X-ray laboratory admin for the
Department of Biophysics and Biophysical
Chemistry of Johns Hopkins University School of Medicine. You can also
read his CV.
We would like to build a community around this website, and we are looking for
volunteers and collaborators to help. Regarding any aspect of the work of this site, please feel free
to contact us,
molecules@gnu-darwin.org, with gdmolecules in the subject line. Cheers!

